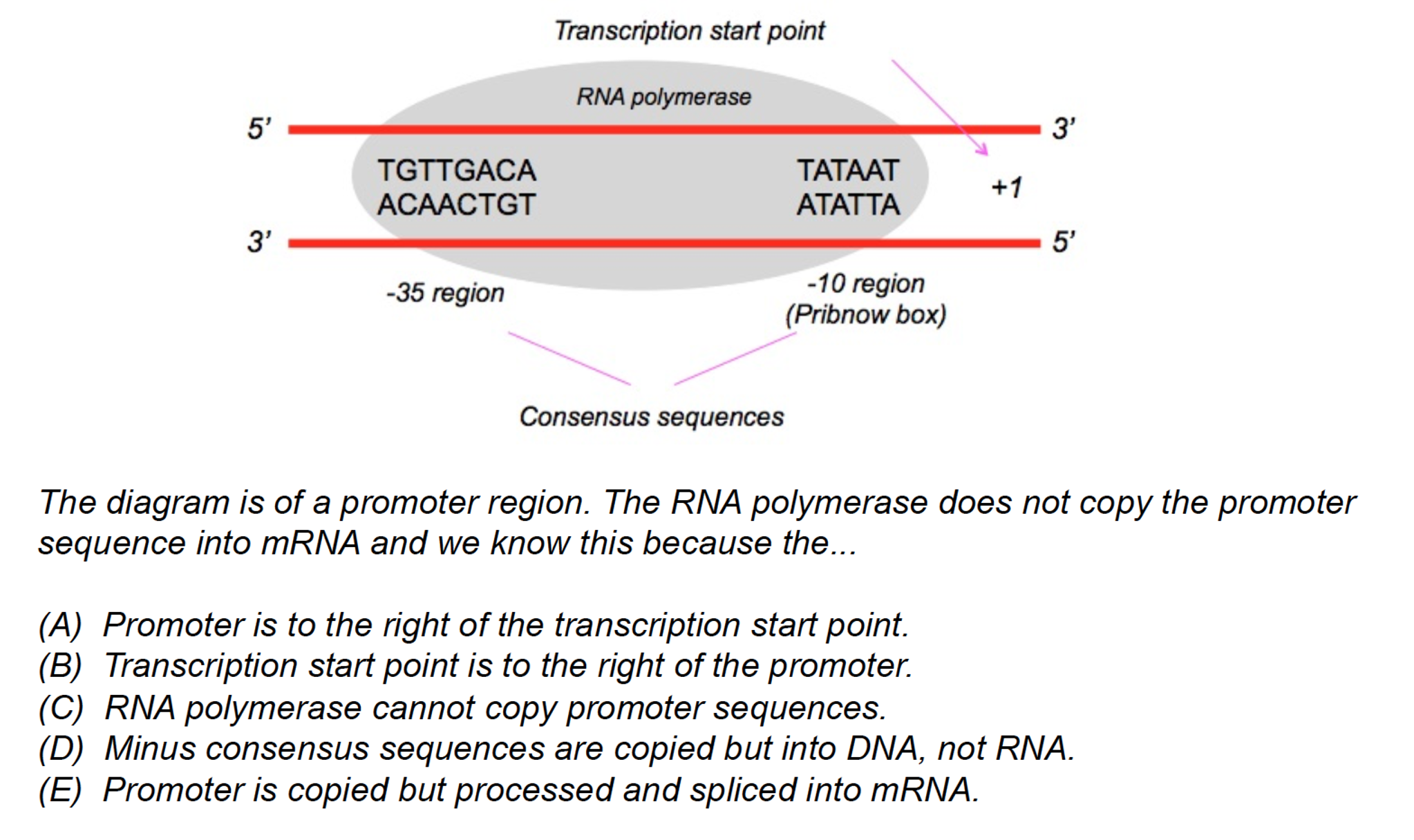
In Escherichia coli, seven alternative sigma factors are responsible for gene expression, while sigma 70 is the most important one as it is required for the expression of housekeeping genes ( 2, 3). While the definition of promoters could vary widely, here we will consider promoters as the core elements recognized by the sigma subunit of the RNAP. Thus, the correct mapping of promoters is a critical step when studying gene expression dynamics in bacteria. Promoter regions are intrinsic DNA elements located upstream of genes and required for their transcription by the RNA polymerase (RNAP) ( 1). We hope the potentials and limitations presented here will help the microbiology community to choose promoter prediction tools among many available alternatives. Using Escherichia coli as a model organism, we demonstrated that while some tools are biased toward AT-rich sequences, others are very efficient in identifying real promoters with low false-negative rates. Here, we assess the predictive power of some of the main prediction tools available using well-defined promoter data sets. Over the last years, many bioinformatics tools have been created to allow users to predict promoter elements in a sequence or genome of interest. Also, when combining new DNA elements into synthetic sequences, predicting the potential generation of new promoter sequences is critical. IMPORTANCE The correct mapping of promoter elements is a crucial step in microbial genomics. 
We present here some potentials and limitations of available tools, and we hope that future work can build upon our effort to systematically characterize this useful class of bioinformatics tools. Of these tools, iPro70-FMWin exhibited the best results for most of the metrics used. We show that the widely used BPROM presented the worse performance among the compared tools, while four tools (CNNProm, iPro70-FMWin, 70ProPred, and iPromoter-2L) offered high predictive power. We compared the performance of the tools using metrics such as specificity, sensitivity, accuracy, and Matthews correlation coefficient (MCC). For this, we used data sets of experimentally validated promoters from Escherichia coli and a control data set composed of randomly generated sequences with similar nucleotide distributions. Here, we performed a systematic comparison between several widely used promoter prediction tools (BPROM, bTSSfinder, BacPP, CNNProm, IBBP, Virtual Footprint, iPro70-FMWin, 70ProPred, iPromoter-2L, and MULTiPly) using well-defined sequence data sets and standardized metrics to determine how well those tools performed related to each other. Additionally, despite many different prediction tools having become popular to identify bacterial promoters, no systematic comparison of such tools has been performed. While new high-throughput technology allows massively parallel mapping of promoter elements, we still mainly rely on bioinformatics tools to predict such elements in bacterial genomes. The promoter region is a key element required for the production of RNA in bacteria.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
